top of page
PUBLICATIONS
Key Publications
* equal contribution. † corresponding author. For the full list of publications, please see Google Scholar.
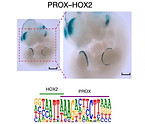
DNA-guided transcription factor interactions extend human gene regulatory code
Xie Z, Sokolov I, Osmala M, Yue X, Bower G, Pett JP, Chen Y, Wang K, Cavga AD, Popov A, Teichmann SA, Morgunova E, Kvon EZ, Yin Y† & Taipale J†.
Nature (2025) PMID40205063 PDF
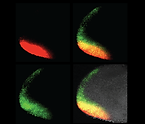
Rapid and Quantitative Functional Interrogation of Human Enhancer Variant Activity in Live Mice
Hollingsworth EW, Liu TA, Alcantara JA, Chen CX, Jacinto SH, Kvon EZ†.
Nature communications (2025) PMID39762235 PDF

Increased Enhancer-Promoter Interactions during Developmental Enhancer Activation in Mammals
Chen Z, Snetkova V, Bower G, Jacinto S, Clock B, Dizehchi A, Barozzi I, Mannion BJ, Alcaina-Caro A, Lopez-Rios J, Dickel DE, Visel A, Pennacchio LA, Kvon EZ†.
Nature Genetics (2024) PMID38509385 PDF DATA

A noncoding single-nucleotide polymorphism at 8q24 drives IDH1-mutant glioma formation
Yanchus C, Drucker KL, Kollmeyer TM, Tsai R, Winick-Ng W, Liang M, Malik A, Pawling J, De Lorenzo SB, Ali A, Decker PA, Kosel ML, Panda A, Al-Zahrani KN, Jiang L, Browning JWL, Lowden C, Geuenich M, Hernandez JJ, Gosio JT, Ahmed M, Loganathan SK, Berman J, Trcka D, Michealraj KA, Fortin J, Carson B, Hollingsworth EW, Jacinto S, Mazrooei P, Zhou L, Elia A, Lupien M, He HH, Murphy DJ, Wang L, Abyzov A, Dennis JW, Maass PG, Campbell K, Wilson MD, Lachance DH, Wrensch M, Wiencke J, Mak T, Pennacchio LA, Dickel DE, Visel A, Wrana J, Taylor MD, Zadeh G, Dirks P, Eckel-Passow JE, Attisano L, Pombo A, Ida CM, Kvon EZ, Jenkins RB, Schramek D.
Science (2022) PMID36201590 PDF

Enhancer redundancy in development and disease
Kvon EZ*†, Waymack R, Elabd MG, Wunderlich Z*†
Nature Reviews Genetics (2021) PMID33442000 PDF

Comprehensive in vivo interrogation reveals phenotypic impact of human enhancer variants
Kvon EZ, Zhu Y, Kelman G, Novak CS, Plajzer-Frick I, Kato M, Garvin TH, Pham Q, Harrington A, Hunter R, Godoy J, Meky EM, Akiyama JA, Afzal V, Tran S, Escande F, Gilbert-Dussardier B, Jean-Marçais N, Hudaiberdiev S, Ovcharenko I, Dobbs MB, Gurnett CA, Manouvrier-Hanu S, Petit F, Visel A, Dickel DE, Pennacchio LA.
Cell (2020) PMID32169219 PDF
Progressive loss of function in a limb enhancer during snake evolution
Kvon EZ, Kamneva OK, Melo US, Barozzi I, Osterwalder M, Mannion BJ, Tissières V, Pickle CS, Plajzer-Frick I, Lee EA, Kato M, Garvin TH, Akiyama JA, Afzal V, Lopez-Rios J, Rubin EM, Dickel DE, Pennacchio LA, Visel A
Cell (2016) PMID27768887 PDF
Using transgenic reporter assays to functionally characterize enhancers in animals
Kvon EZ†
Genomics (2015) PMID26072435 PDF
Genome-scale functional characterization of Drosophila developmental enhancers in vivo
Kvon EZ, Kazmar T, Stampfel G, Yanez-Cuna JO, Pagani M, Schernhuber K, Dickson BJ, Stark A
Nature (2014) PMID24896182 PDF
Deciphering the transcriptional cis-regulatory code
Yanez-Cuna JO*, Kvon EZ*, Stark A
Trends in Genetics (2013) PMID23102583 PDF
HOT regions function as patterned developmental enhancers and have a distinct cis-regulatory signature
Kvon EZ*, Stampfel G.*, Yáñez-Cuna JO, Dickson, BJ, Stark, A
Genes & Development (2012) PMID22499593 PDF

Genetic Factors Mediating Long-range Enhancer–Promoter Communication in Mammalian Development
Bower G, Kvon EZ†.
Current Opinion in Genetics & Development (2024) PMID39579740 PDF

The Genomes of All Lungfish Inform on Genome Expansion and Tetrapod Evolution
BSchartl M†, Woltering JM, Irisarri I, Du K, Kneitz S, Pippel M, Brown T, Franchini P, Li J, Li M, Adolfi M, Winkler S, Sousa JF, Chen Z, Jacinto S, Kvon EZ, Oliveira LRC, Monteiro E, Amaral DB, Burmester T, Chalopin D, Suh A, Myers E, Simakov O, Schneider I, Meyer A†.
Nature (2024) PDF
_edited.jpg)
Emx2 Underlies the Development and Evolution of Marsupial Gliding Membranes
Moreno JA, Dudchenko O, YF Charles, Mereby SA, Chen Z, Ramos R, Almet AA, Sen H, Brack BJ, Johnson MR, Li S, Wang W, Gaska JM, Ploss A, Weisz D, Omer AD, Yao W, Colaric Z, Kaur P, Leger JS, Nie Q, Mena A, Flanagan JP, Keller G, Sanger T, Ostrow B, Plikus MV, Kvon EZ, Aiden EL† & Mallarino R†.
Nature (2024) PMC11062917 PDF
bottom of page